Answer:
DNA FINGER PRINTING
Definition. Technique of using
DNA fragments, resulting from restriction endonuclease enzyme cleavage to
identify particular individuals, is called DNA finger printing.
Introduction. DNA finger
printing technique was developed by Alee Jffery (1985, 86) at Leicester
University, United Kingdom.
Inheritance of DNA is very
stable. Every person has specific pattern of DNA sequence which shows a
combination of DNA sequence of both mother and father. Study of DNA finger
prints helps in establishment of identity of a person, identification of
criminals in case of a murder or rape, and paternity test in case of disputed parentage.
This technique helps in identification even when the stains on victims clothes
etc. are even several years old.
In India, first test of DNA
finger printing was done in June, 1989 for setting a disputed parentage in
Madras. Laboratory for DNA finger printing is situated in Hyderabad at the
Centre for Cell and Molecular Biology (CCMB). Paternity dispute cases are much
more common in India and most of them are referred to CCMB for DNA evidence.
Southern Blotting technique of
DNA finger printing:
VNTR (Variable number of tandem
repeats) technique involved Southern blot hybridization using radiolabelled
VNTR as a probe. The steps were:
(a) For DNA finger printing, DNA
is isolated from anybody cell or even from blood stains, semen stains or hair
roots.
(b) DNA sample is first digested
with a restriction endonuclease enzyme and the digested sample is then gel
electrophoresed.
(c) DNA bonds in the gel are
denatured into single strands by treating them with alkali solution.
(d) Gel is laid on the top of a
buffer saturated filter paper, placed on a solid support (e.g. glass plate),
with its two edges immersed in the buffer.
(e) A sheet of nitrocellulose
membrane is placed on top of the gel and a stack of paper towels is placed on
the top of this membrane (acts as an absorbent tissue).
(f) A weight of about 0.5 kg is
placed on the top of pack of paper towels. Leave this arrangement for a few
hours or overnight.
The buffer solution is drawn up
by the filter paper wick, and passes through the gel to the nitrocellulose
membrane and finally to the paper towels. While passing through the gel, the
buffer carries with it single stranded DNA molecules, which bind to the
nitrocellulose membrane, while the buffer passes through it to the paper towels.
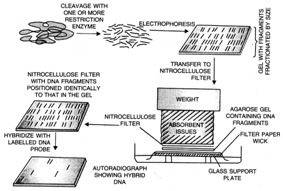 Soluthern blotting
technique of DNA finger printing.
(g) Next day, paper towels are
removed and discarded.
(h) Nitrocellulose membrane with
single stranded DNA bands is incubated at 80°C for the 2-3 hours to fix the DNA
bands permanently on the membrane.
(i) This membrane now has a
replica of DNA bands from agarose gel and can be used for hybridization with
specific labelled DNA or RNA probes.
(j) The membrane may then be
washed to remove any unwound DNA and X-ray film is exposed to the hybridized
membrane to get autoradiograph.
Significance:
DNA finger printing shows
polymorphism of DNA which is used for identification of a personous with much
more certainty that has hitherto been possible through techniques of blood
groups since the number of blood groups available becomes a limitation. The technique
has shown the possibility of two persons having same pattern of DNA finger
prints is very remote (exception monozygotic i.e., identical twin.)
? The number of repeats m
VNTR shows high degree of polymorphism.
? The size of VNTR differs
from 0.1 to 20 kb. In autoradiogram after hybridization with VNTR probe several
bands of varying sizes appear.
? Alec Jeffreys used
satellite DNA as probe that indicated high degree of polymorphism and was
called as VNTR (variable number of tandem repeats).
? VNTR belongs to a class of
satellite DNA called as mini-satellite.
? PCR (polymerase chain
reaction increases the sensitivity of technique).
? If an inheritable mutation
4s observed in a population of high frequency, it is called as DNA
polymorphism.
Soluthern blotting
technique of DNA finger printing.
(g) Next day, paper towels are
removed and discarded.
(h) Nitrocellulose membrane with
single stranded DNA bands is incubated at 80°C for the 2-3 hours to fix the DNA
bands permanently on the membrane.
(i) This membrane now has a
replica of DNA bands from agarose gel and can be used for hybridization with
specific labelled DNA or RNA probes.
(j) The membrane may then be
washed to remove any unwound DNA and X-ray film is exposed to the hybridized
membrane to get autoradiograph.
Significance:
DNA finger printing shows
polymorphism of DNA which is used for identification of a personous with much
more certainty that has hitherto been possible through techniques of blood
groups since the number of blood groups available becomes a limitation. The technique
has shown the possibility of two persons having same pattern of DNA finger
prints is very remote (exception monozygotic i.e., identical twin.)
? The number of repeats m
VNTR shows high degree of polymorphism.
? The size of VNTR differs
from 0.1 to 20 kb. In autoradiogram after hybridization with VNTR probe several
bands of varying sizes appear.
? Alec Jeffreys used
satellite DNA as probe that indicated high degree of polymorphism and was
called as VNTR (variable number of tandem repeats).
? VNTR belongs to a class of
satellite DNA called as mini-satellite.
? PCR (polymerase chain
reaction increases the sensitivity of technique).
? If an inheritable mutation
4s observed in a population of high frequency, it is called as DNA
polymorphism.
You need to login to perform this action.
You will be redirected in
3 sec
